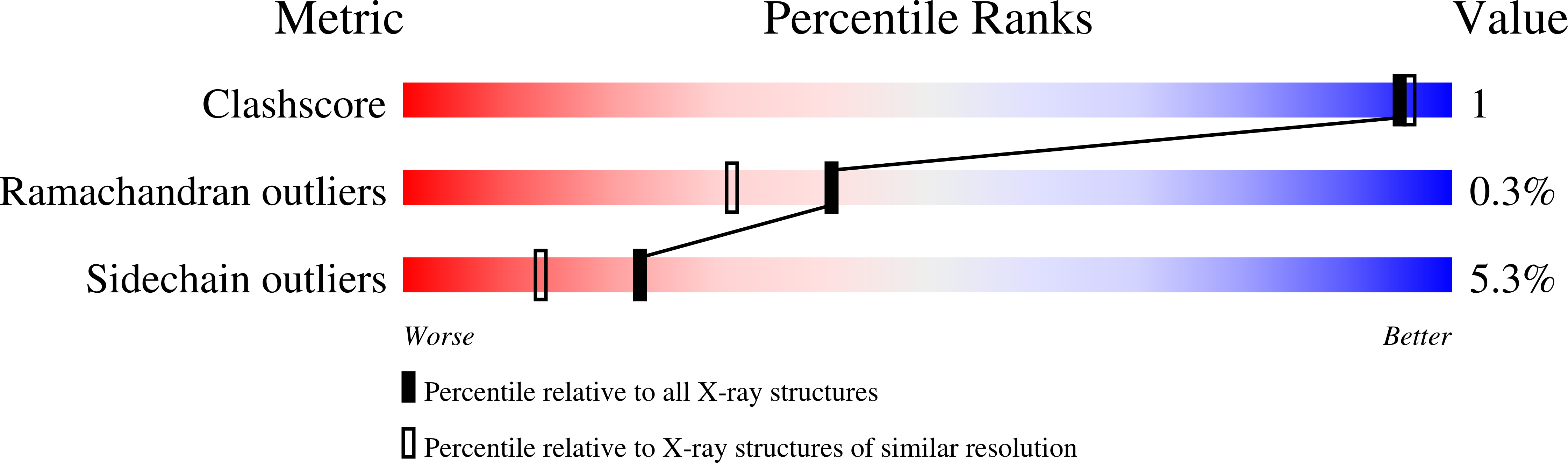
RCSB PDB - 8XIA: X-RAY ANALYSIS OF D-XYLOSE ISOMERASE AT 1.9 ANGSTROMS: NATIVE ENZYME IN COMPLEX WITH SUBSTRATE AND WITH A MECHANISM-DESIGNED INACTIVATOR

Applied Sciences | Free Full-Text | Glucose Isomerase: Functions, Structures, and Applications | HTML

Locating active-site hydrogen atoms in d-xylose isomerase: Time-of-flight neutron diffraction | PNAS

In vivo and in vitro reactions catalyzed by xylose isomerase (XI) and AI | Download Scientific Diagram
![PDF] Metal ion roles and the movement of hydrogen during reaction catalyzed by D-xylose isomerase: a joint x-ray and neutron diffraction study. | Semantic Scholar PDF] Metal ion roles and the movement of hydrogen during reaction catalyzed by D-xylose isomerase: a joint x-ray and neutron diffraction study. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/69ce4fdb8daad44b9853bda2a4c50e3df9acb402/2-Figure1-1.png)
PDF] Metal ion roles and the movement of hydrogen during reaction catalyzed by D-xylose isomerase: a joint x-ray and neutron diffraction study. | Semantic Scholar

Probing the Roles of Active Site Residues in D-Xylose Isomerase (∗) - Journal of Biological Chemistry

Catalytic reaction mechanism of Pseudomonas stutzeri l‐rhamnose isomerase deduced from X‐ray structures - Yoshida - 2010 - The FEBS Journal - Wiley Online Library

Figure 9 from Catalytic mechanism of xylose (glucose) isomerase from Clostridium thermosulfurogenes. Characterization of the structural gene and function of active site histidine. | Semantic Scholar

Xylose fermentation efficiency of industrial Saccharomyces cerevisiae yeast with separate or combined xylose reductase/xylitol dehydrogenase and xylose isomerase pathways | Biotechnology for Biofuels and Bioproducts | Full Text

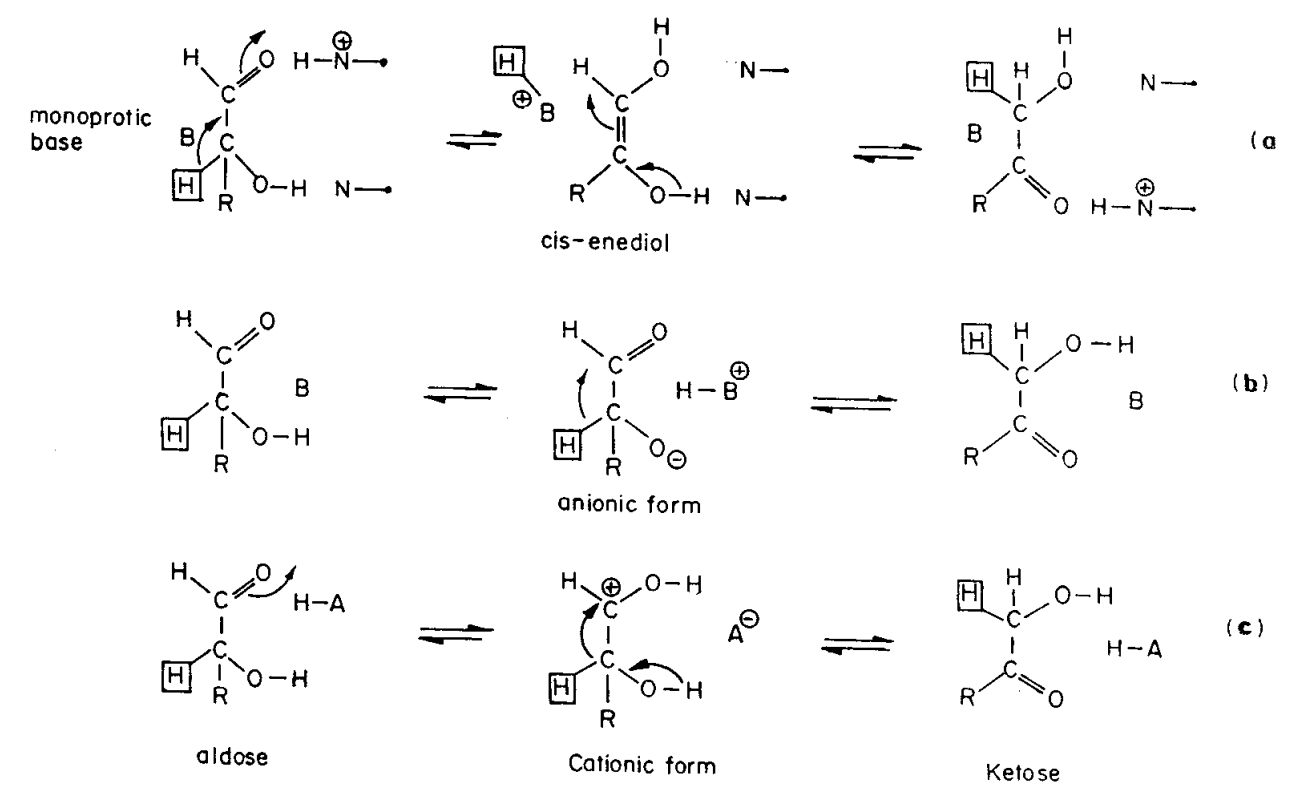
Catalytic mechanism of xylose isomerase. Top: ring opening, middle:... | Download Scientific Diagram

L-Arabinose Binding, Isomerization, and Epimerization by D-Xylose Isomerase: X-Ray/Neutron Crystallographic and Molecular Simulation Study: Structure

Structural insight into D-xylose utilization by xylose reductase from Scheffersomyces stipitis | Scientific Reports
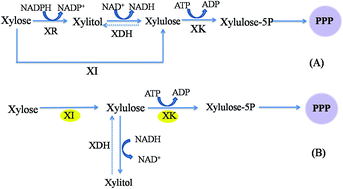
The role of a xylose isomerase pathway in the conversion of xylose to lipid in Mucor circinelloides - RSC Advances (RSC Publishing)
Proposed catalytic reaction mechanism including sugar-ring openings of... | Download Scientific Diagram
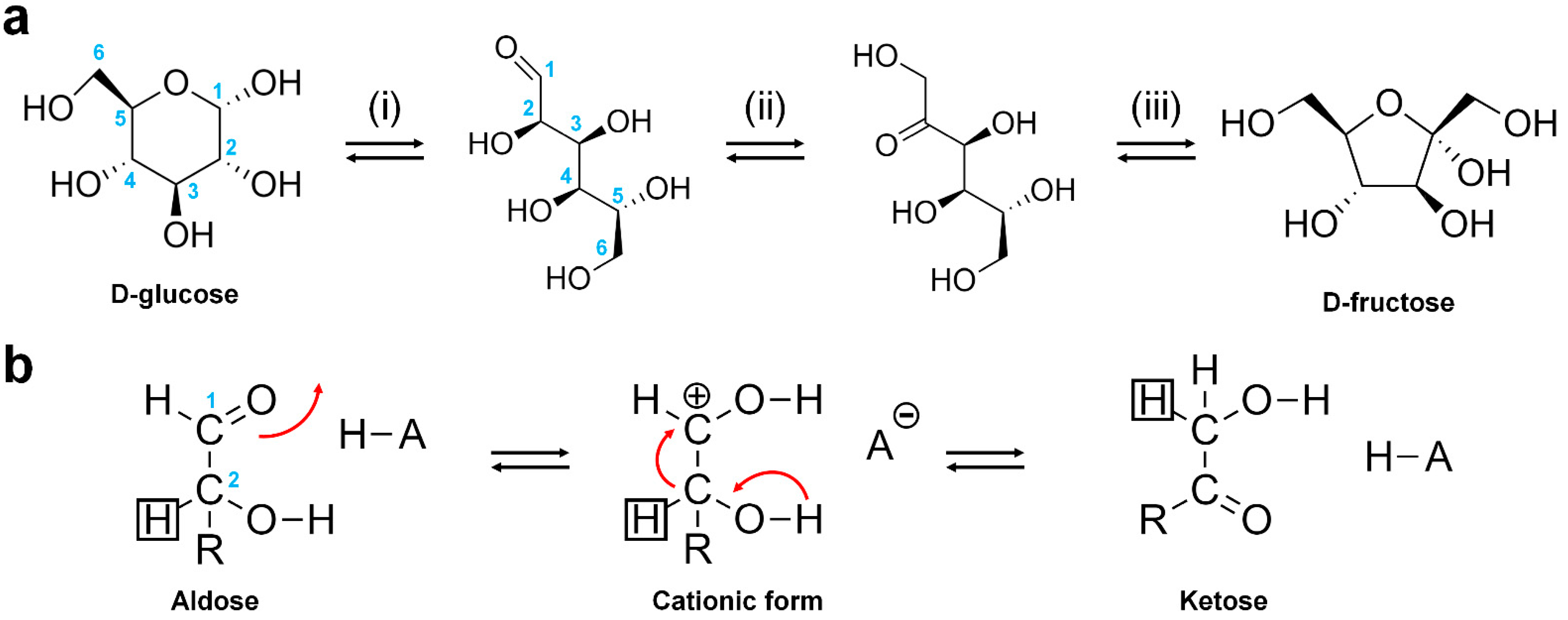
Applied Sciences | Free Full-Text | Glucose Isomerase: Functions, Structures, and Applications | HTML

Magnesium in PDB 2xis: A Metal-Mediated Hydride Shift Mechanism For Xylose Isomerase Based on the 1.6 Angstroms Streptomyces Rubiginosus Structures with Xylitol and D-Xylose