T7 RNA polymerase‐driven inducible cell lysis for DNA transfer from Escherichia coli to Bacillus subtilis - Juhas - 2017 - Microbial Biotechnology - Wiley Online Library

A novel molecular beacon-based method for isothermal detection of sequence-specific DNA via T7 RNA polymerase-aided target regeneration - ScienceDirect

A single mutation attenuates both the transcription termination and RNA-dependent RNA polymerase activity of T7 RNA polymerase | bioRxiv

Isothermal detection of lncRNA using T7 RNA polymerase mediated amplification coupled with fluorescence-based sensor - ScienceDirect
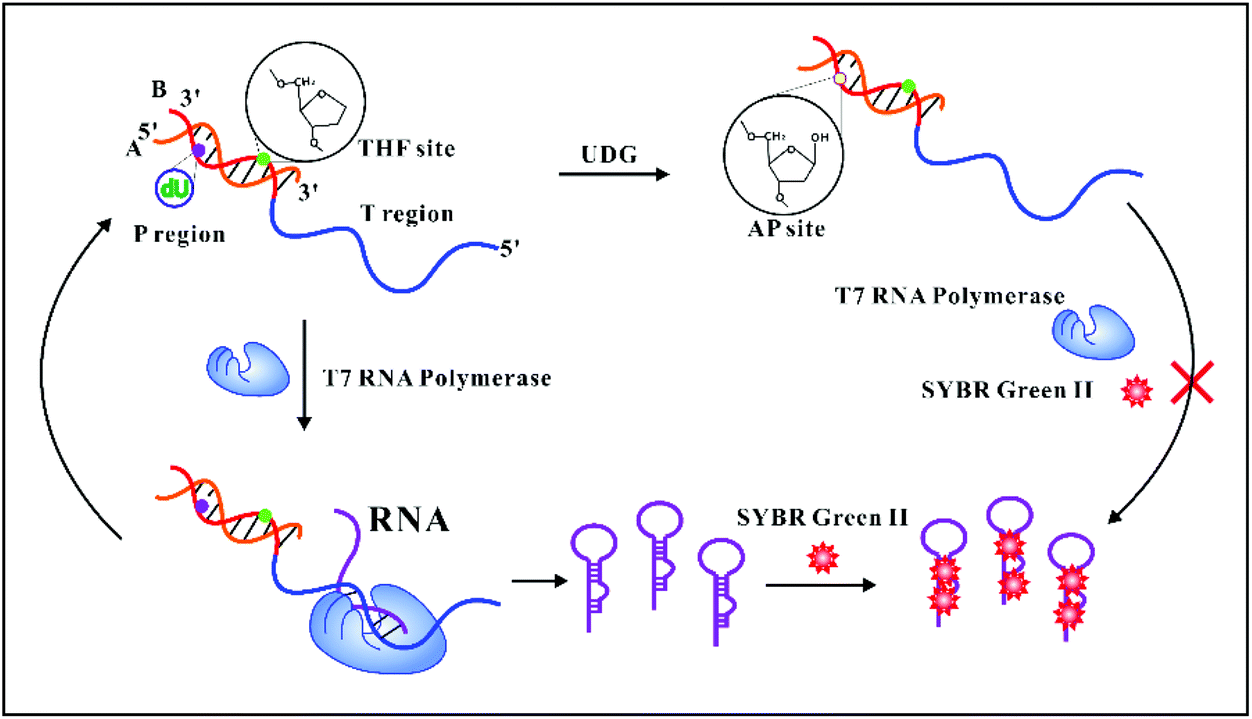
A novel analytical principle using AP site-mediated T7 RNA polymerase transcription regulation for sensing uracil-DNA glycosylase activity - Analyst (RSC Publishing) DOI:10.1039/D0AN00509F
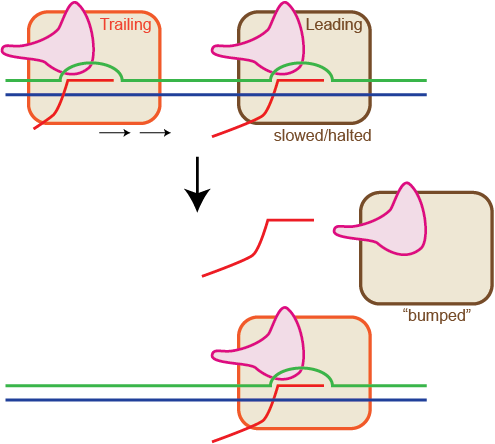
Observed instability of T7 RNA polymerase elongation complexes can be dominated by collision-induced “bumping” – Martin Lab
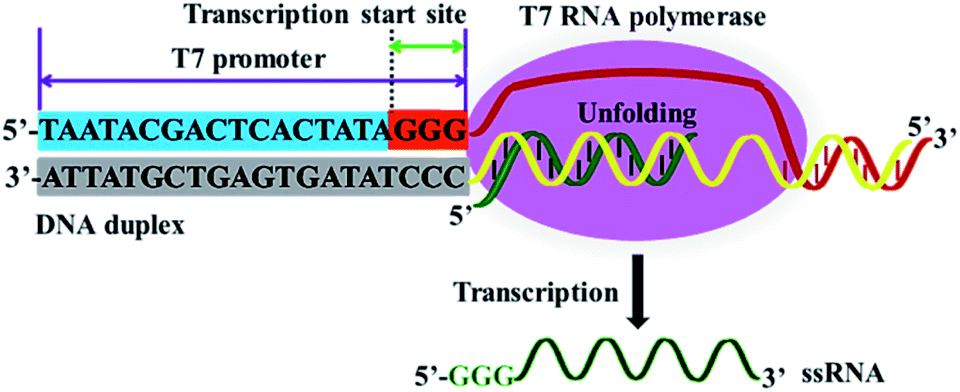
A controlled T7 transcription-driven symmetric amplification cascade machinery for single-molecule detection of multiple repair glycosylases - Chemical Science (RSC Publishing) DOI:10.1039/D1SC00189B

In vivo diversification of target genomic sites using processive T7 RNA polymerase-base deaminase fusions blocked by RNA-guided dCas9 | bioRxiv
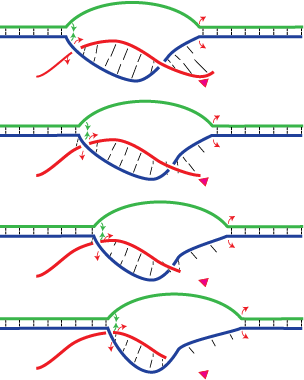
Dissociation of halted T7 RNA polymerase elongation complexes proceeds via a forward translocation mechanism – Martin Lab
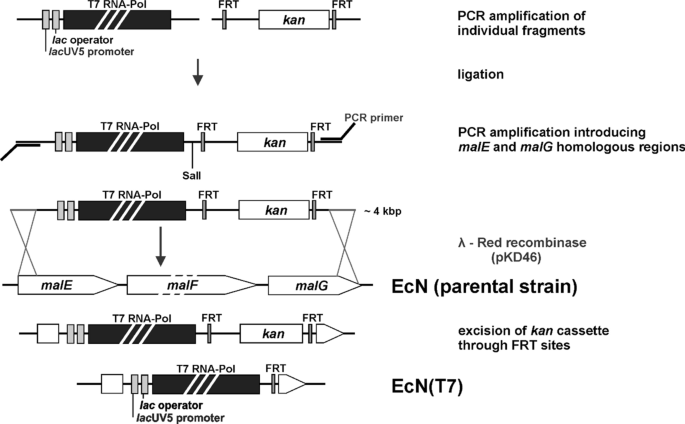
Construction of a new T7 promoter compatible Escherichia coli Nissle 1917 strain for recombinant production of heme-dependent proteins | Microbial Cell Factories | Full Text

The specificity loop of T7 RNA polymerase interacts first with the promoter and then with the elongating transcript, suggesting a mechanism for promoter clearance | PNAS

Dissociation of halted T7 RNA polymerase elongation complexes proceeds via a forward-translocation mechanism | PNAS
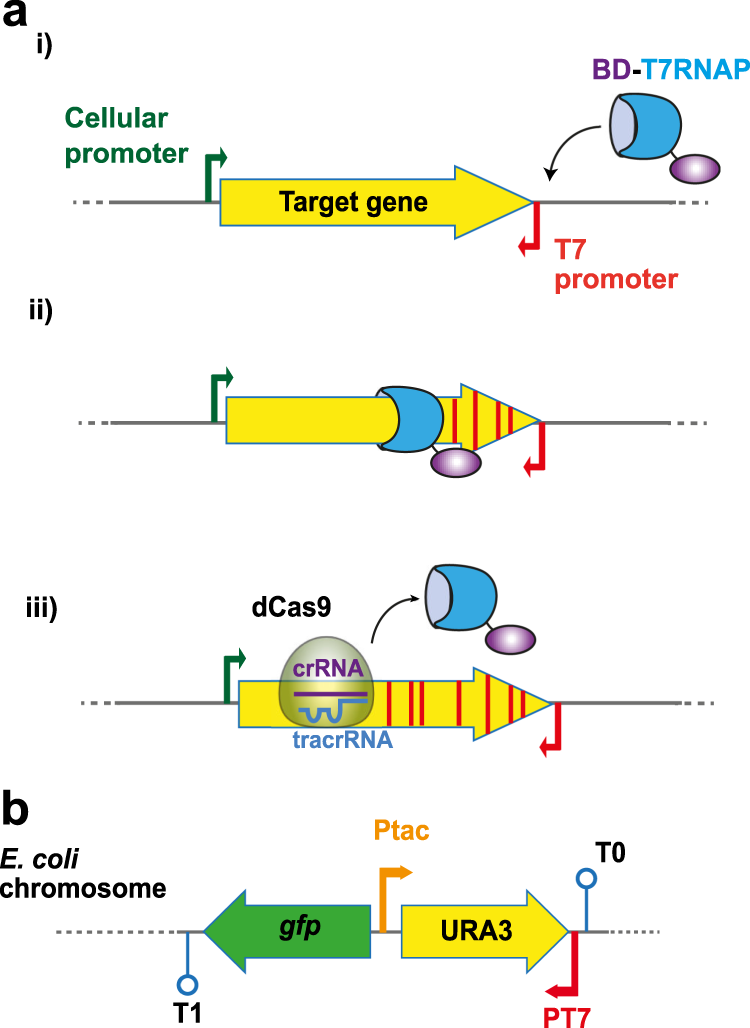
In vivo diversification of target genomic sites using processive base deaminase fusions blocked by dCas9 | Nature Communications

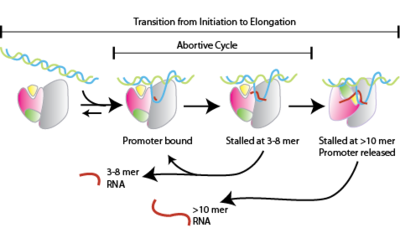
Mechanism of Instability in Abortive Cycling by T7 RNA Polymerase* - Journal of Biological Chemistry