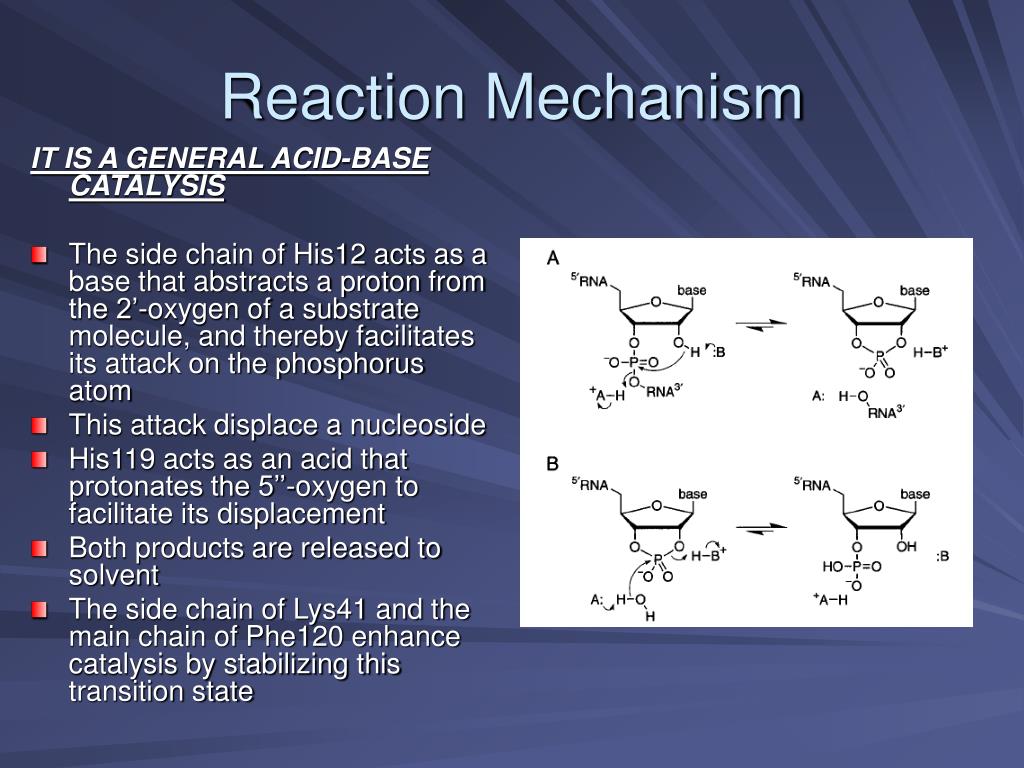
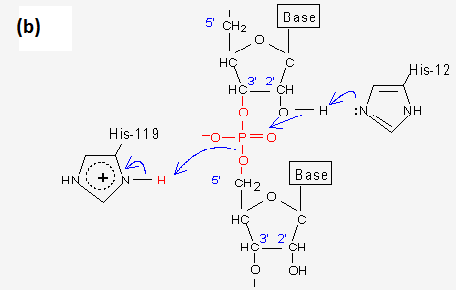
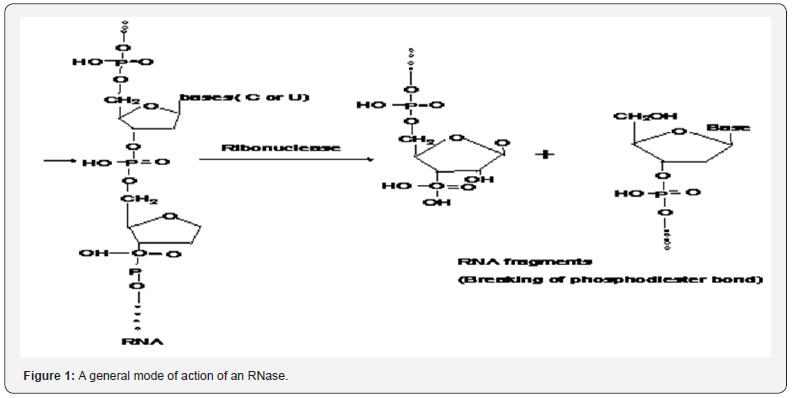
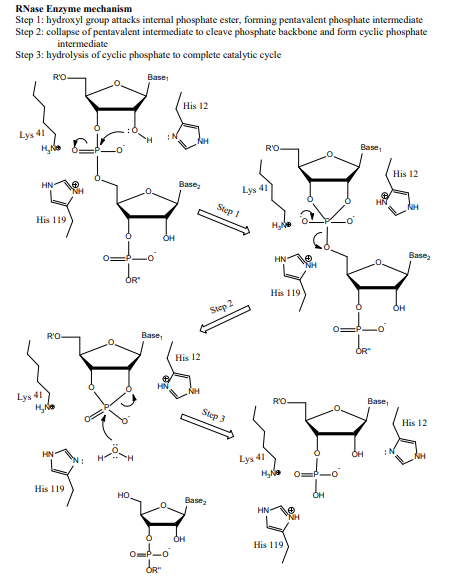
Mechanism of RNase T1: concerted triester-like phosphoryl transfer via a catalytic three-centered hydrogen bond - ScienceDirect
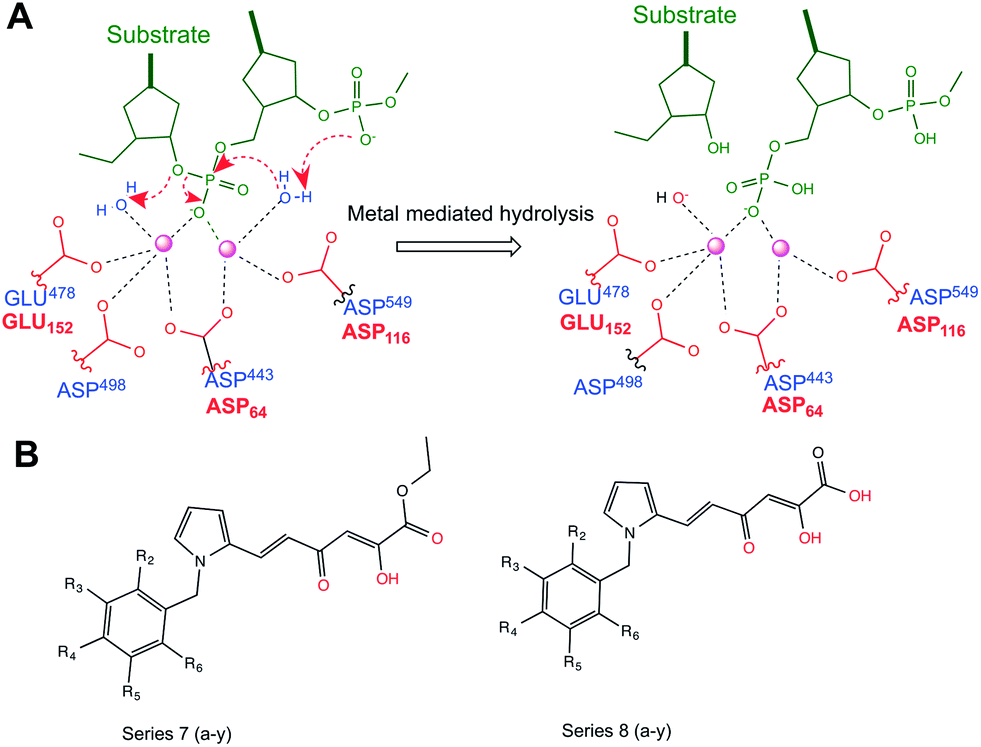
Dual mechanism of HIV-1 integrase and RNase H inhibition by diketo derivatives – a computational study - RSC Advances (RSC Publishing) DOI:10.1039/C4RA05728G
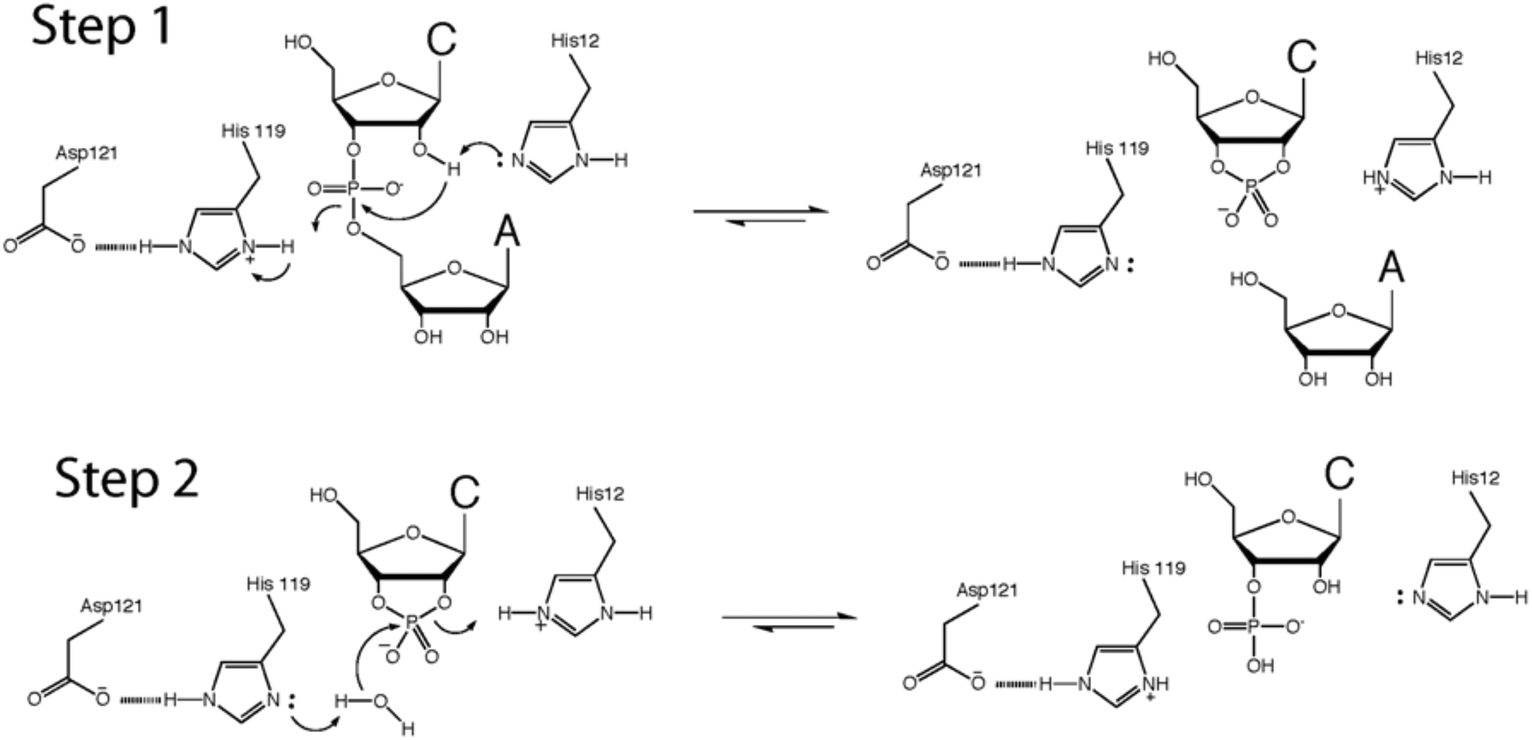
Chemical synthesis and enzymatic properties of RNase A analogues designed to enhance second-step catalytic activity - Organic & Biomolecular Chemistry (RSC Publishing) DOI:10.1039/C6OB01163B

Mitochondrial ribonuclease P structure provides insight into the evolution of catalytic strategies for precursor-tRNA 5′ processing | PNAS

The Subsites Structure of Bovine Pancreatic Ribonuclease A Accounts for the Abnormal Kinetic Behavior with Cytidine 2′,3′-Cyclic Phosphate* - Journal of Biological Chemistry
PLOS ONE: Antisense Oligonucleotides Capable of Promoting Specific Target mRNA Reduction via Competing RNase H1-Dependent and Independent Mechanisms