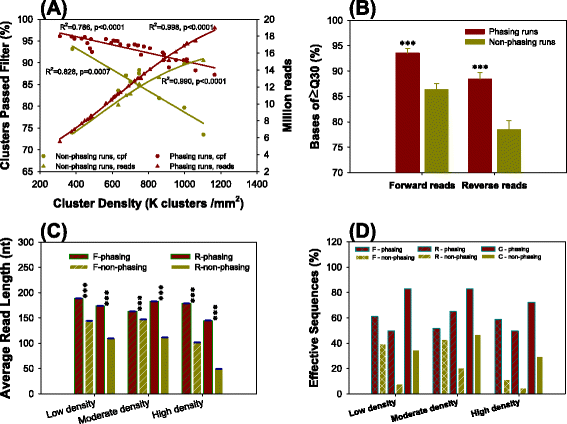
In‐depth comparative analysis of Illumina® MiSeq run metrics: Development of a wet‐lab quality assessment tool - Kastanis - 2019 - Molecular Ecology Resources - Wiley Online Library

In‐depth comparative analysis of Illumina® MiSeq run metrics: Development of a wet‐lab quality assessment tool - Kastanis - 2019 - Molecular Ecology Resources - Wiley Online Library

Illumina iSeq 100 and MiSeq exhibit similar performance in freshwater fish environmental DNA metabarcoding | Scientific Reports

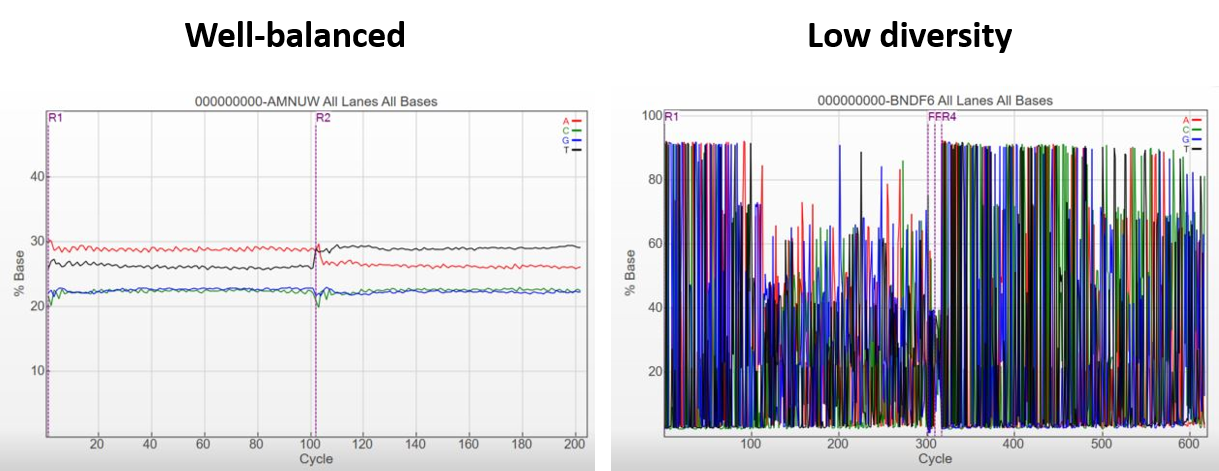
Plotting % Occupied by % Pass Filter to optimize loading concentration for NovaSeq and iSeq 100 platforms

In‐depth comparative analysis of Illumina® MiSeq run metrics: Development of a wet‐lab quality assessment tool - Kastanis - 2019 - Molecular Ecology Resources - Wiley Online Library
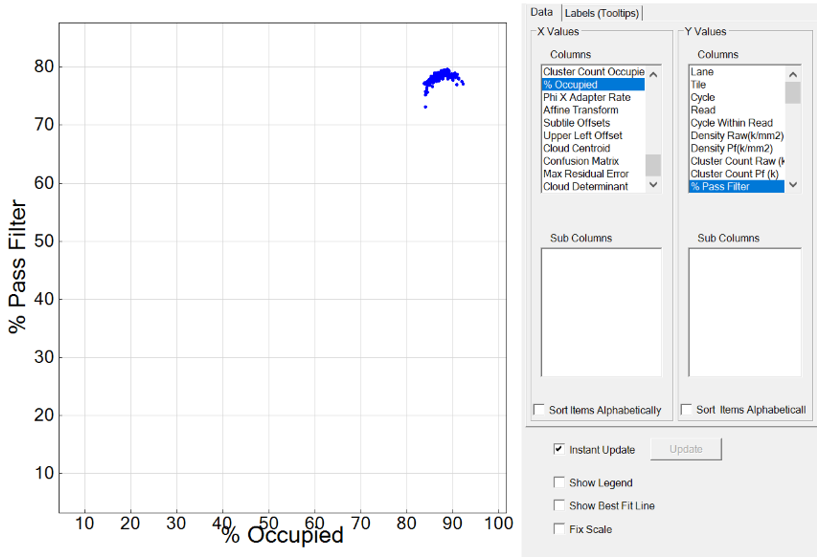
Plotting % Occupied by % Pass Filter to optimize loading concentration for NovaSeq and iSeq 100 platforms

A 16S rRNA gene sequencing and analysis protocol for the Illumina MiniSeq platform - Pichler - 2018 - MicrobiologyOpen - Wiley Online Library

gDNA Enrichment by a Transposase-based Technology for NGS Analysis of the Whole Sequence of BRCA1, BRCA2, and 9 Genes Involved in DNA Damage Repair | Protocol

Clusters Passing Filter‚’œ€é©Œ–™‚‹«¯ PF = (x100)* 11 ©†™‚Œ°%Clusters Passing Filter‚’祂‹“¨ ...