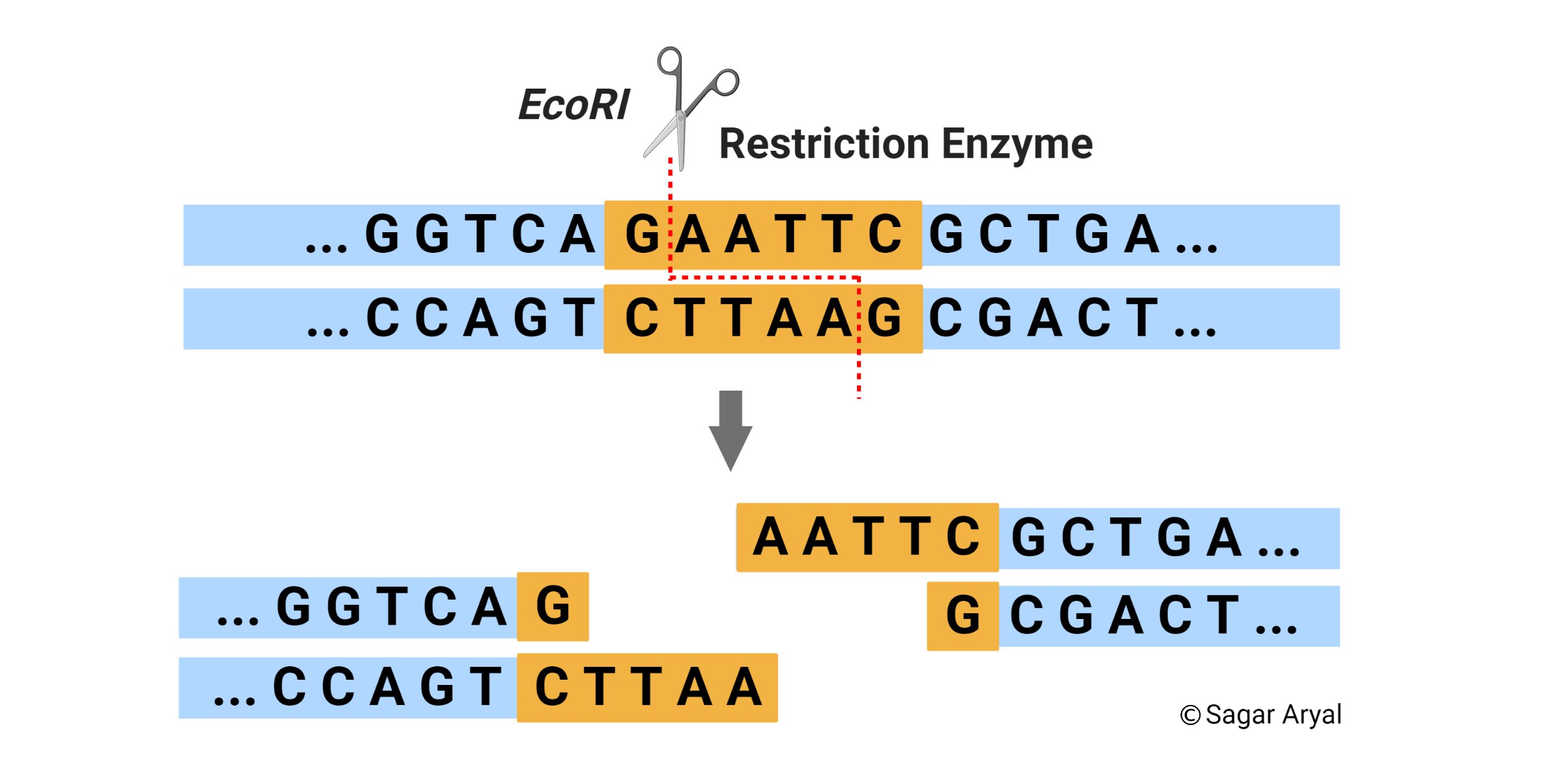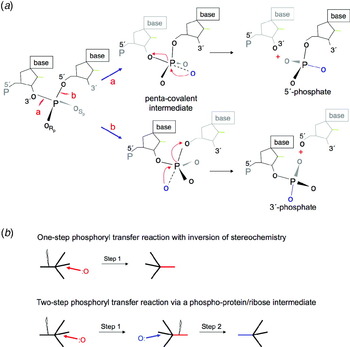
Nucleases: diversity of structure, function and mechanism | Quarterly Reviews of Biophysics | Cambridge Core

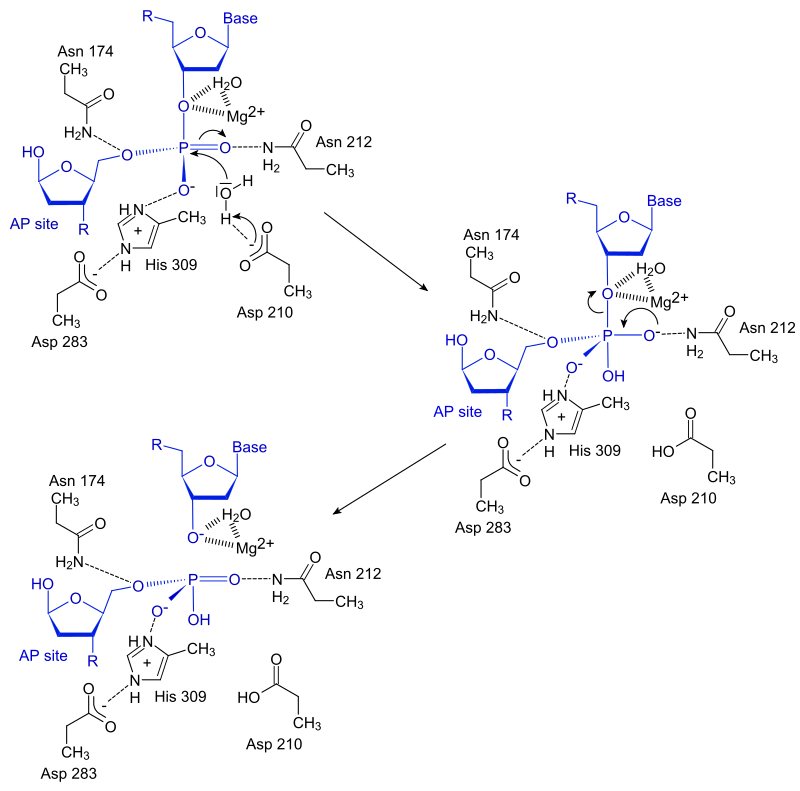
Proposed three-metal ion mechanism for EcoRV endonuclease . Reprinted... | Download Scientific Diagram

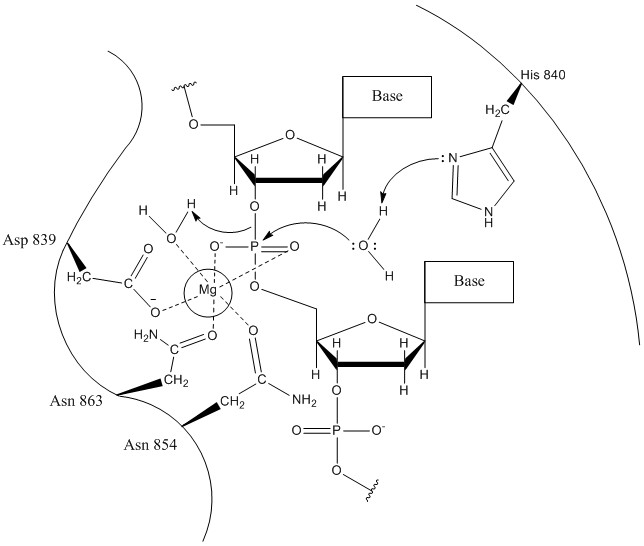
Involvement of Glutamic Acid 23 in the Catalytic Mechanism of T4 Endonuclease V (∗) - Journal of Biological Chemistry

Proposed three-metal ion mechanism for EcoRV endonuclease . Reprinted... | Download Scientific Diagram
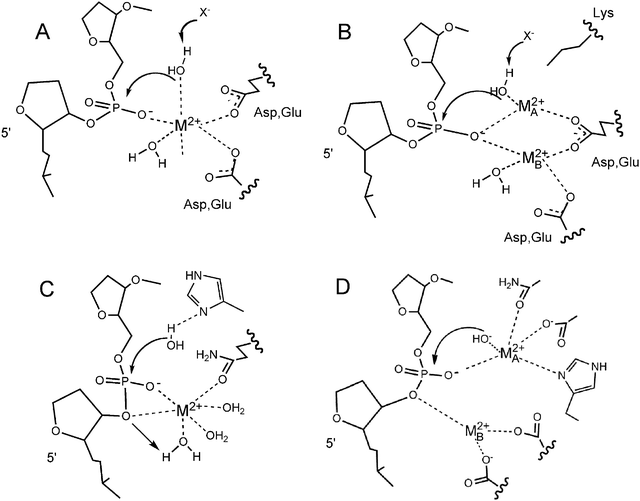
One is enough: insights into the two-metal ion nuclease mechanism from global analysis and computational studies - Metallomics (RSC Publishing) DOI:10.1039/C0MT00013B
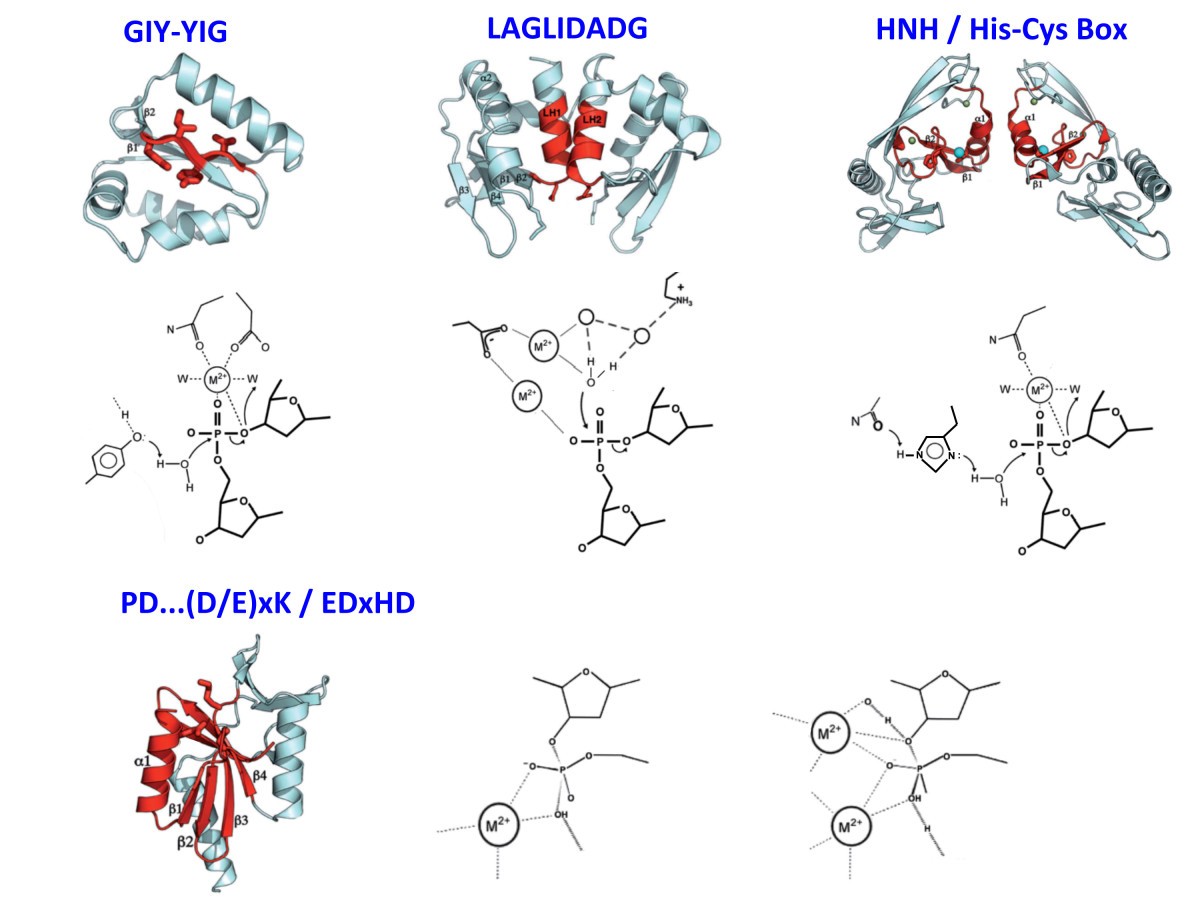
Homing endonucleases from mobile group I introns: discovery to genome engineering | Mobile DNA | Full Text
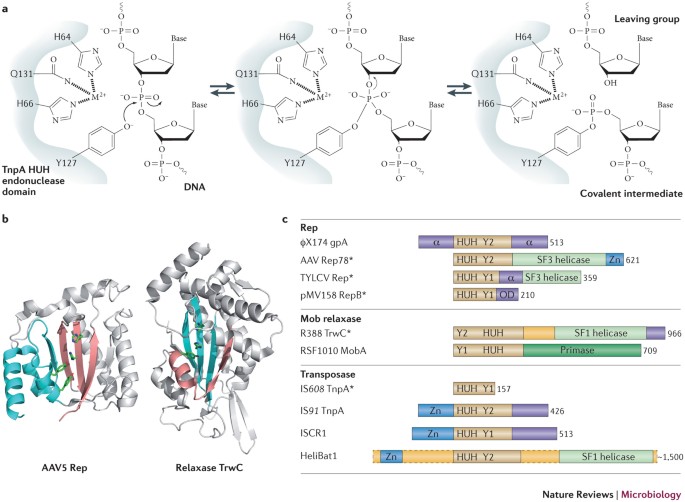
Breaking and joining single-stranded DNA: the HUH endonuclease superfamily | Nature Reviews Microbiology

Figure 3 from Catalytic mechanism of DNA backbone cleavage by the restriction enzyme EcoRV: a quantum mechanical/molecular mechanical analysis. | Semantic Scholar

Structural Determinants for Specific Recognition by T4 Endonuclease V* - Journal of Biological Chemistry
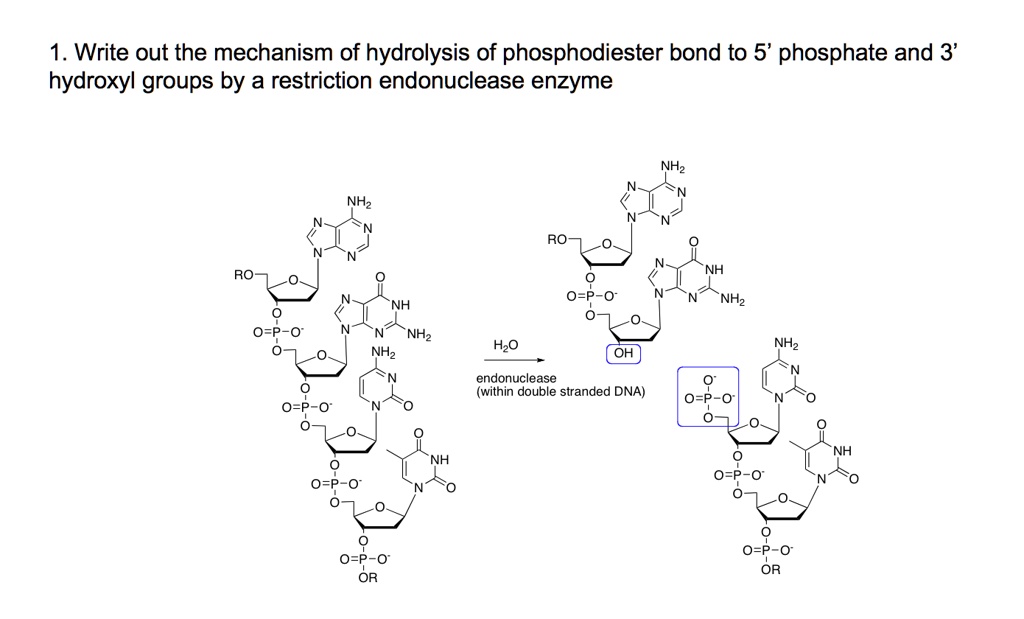
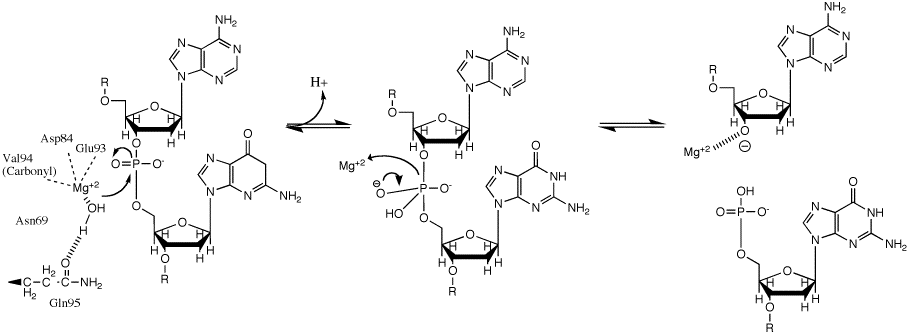
SOLVED:1 . Write out the mechanism of hydrolysis of phosphodiester bond to 5' phosphate and 3' hydroxyl groups by a restriction endonuclease enzyme NHz NHz RO RO NH NHz NHz NHz Hzo